Python实现GB格式序列文件转换Fasta格式文件
目录
- GB格式文件和FASTA文件介绍
- 处理步骤
- Python脚本代码如下:
- 运行情况
GB格式文件和FASTA文件介绍
在分子生物学中 我们会有将GB格式序列文件 转换成 Fasta格式文件的需求,这里我们利用python脚本来解决这个问题。
gb格式文件是GenBank的文件,用来保存序列的详细信息。包含一个gene的名称,编号,发现者,参考文献,外显子位置,编码区序列,蛋白序列等等信息。
例如:
LOCUS NM_213806 849 bp mRNA linear MAM 24-SEP-2019
DEFINITION Sus scrofa Fas ligand (TNF superfamily, member 6) (FASLG), mRNA.
ACCESSION NM_213806
VERSION NM_213806.1
KEYWORDS RefSeq.
SOURCE Sus scrofa (pig)
ORGANISM Sus scrofa
Eukaryota; Metazoa; Chordata; Craniata; Vertebrata; Euteleostomi;
Mammalia; Eutheria; Laurasiatheria; Cetartiodactyla; Suina; Suidae;
Sus.
REFERENCE 1 (bases 1 to 849)
AUTHORS Lin F, Fu YH, Han J, Shen M, Du CW, Li R, Ma XS and Liu HL.
TITLE Changes in the expression of Fox O1 and death ligand genes during
follicular atresia in porcine ovary
JOURNAL Genet. Mol. Res. 13 (3), 6638-6645 (2014)
PUBMED 25177944
REMARK GeneRIF: Data suggest forkhead box protein O1 (FoxO1) involvement
in the regulation of TNF-related apoptosis-inducing ligand TRAIL
and Fas ligand FasL expression during follicular atresia.
Publication Status: Online-Only
REFERENCE 2 (bases 1 to 849)
AUTHORS Xie GH, Wang SJ, Wang Y, Zhang Y, Zhang HZ, Jin S, Wang QF, Liu ZC
and Ge HL.
TITLE Fas Ligand gene transfer enhances the survival of tissue-engineered
chondrocyte allografts in mini-pigs
JOURNAL Transpl. Immunol. 19 (2), 145-151 (2008)
PUBMED 18503890
REMARK GeneRIF: the result indicates that the expression of FasL by
chondrocytes is capable of inducing apoptosis of activated T cells
REFERENCE 3 (bases 1 to 849)
AUTHORS Chang HW, Jeng CR, Lin CM, Liu JJ, Chang CC, Tsai YC, Chia MY and
Pang VF.
TITLE The involvement of Fas/FasL interaction in porcine circovirus type
2 and porcine reproductive and respiratory syndrome virus
co-inoculation-associated lymphocyte apoptosis in vitro
JOURNAL Vet. Microbiol. 122 (1-2), 72-82 (2007)
PUBMED 17321702
REMARK GeneRIF: The expression of FAS and FAS ligand in splenic
macrophages co-infected with porcine circovirus 2 and porcine
reproductive and respiratory syndrome virus is reported
REFERENCE 4 (bases 1 to 849)
AUTHORS Tayade C, Black GP, Fang Y and Croy BA.
TITLE Differential gene expression in endometrium, endometrial
lymphocytes, and trophoblasts during successful and abortive embryo
implantation
JOURNAL J. Immunol. 176 (1), 148-156 (2006)
PUBMED 16365405
REFERENCE 5 (bases 1 to 849)
AUTHORS Bai L, Maedler K, Donath M and Tuch BE.
TITLE Expression of Fas but not Fas ligand on fetal pig beta cells
JOURNAL Xenotransplantation 11 (5), 426-435 (2004)
PUBMED 15303979
REMARK GeneRIF: FasL was not detected on fetal pig pancreatic cells but
could be induced on both beta and non-beta cells when the cells
were treated with IL1beta.
Erratum:[Xenotransplantation. 2016 Mar;23(2):171-2. PMID: 27106874]
REFERENCE 6 (bases 1 to 849)
AUTHORS Tsuyuki S, Kono M and Bloom ET.
TITLE Cloning and potential utility of porcine Fas ligand: overexpression
in porcine endothelial cells protects them from attack by human
cytolytic cells
JOURNAL Xenotransplantation 9 (6), 410-421 (2002)
PUBMED 12371937
REFERENCE 7 (bases 1 to 849)
AUTHORS Motegi-Ishiyama Y, Nakajima Y, Hoka S and Takagaki Y.
TITLE Porcine Fas-ligand gene: genomic sequence analysis and comparison
with human gene
JOURNAL Mol. Immunol. 38 (8), 581-586 (2002)
PUBMED 11792426
REFERENCE 8 (bases 1 to 849)
AUTHORS Muneta Y, Shimoji Y, Inumaru S and Mori Y.
TITLE Molecular cloning, characterization, and expression of porcine Fas
ligand (CD95 ligand)
JOURNAL J. Interferon Cytokine Res. 21 (5), 305-312 (2001)
PUBMED 11429161
COMMENT PROVISIONAL REFSEQ: This record has not yet been subject to final
NCBI review. The reference sequence was derived from AB027297.1.
##Evidence-Data-START##
Transcript exon combination :: AB027297.1, AF397407.1 [ECO:0000332]
RNAseq introns :: single sample supports all introns
SAMN01893940, SAMN01915393
[ECO:0000348]
##Evidence-Data-END##
FEATURES Location/Qualifiers
source 1..849
/organism="Sus scrofa"
/mol_type="mRNA"
/db_xref="taxon:9823"
/chromosome="9"
/map="9"
gene 1..849
/gene="FASLG"
/gene_synonym="CD95-L; FASL; TNFSF6"
/note="Fas ligand (TNF superfamily, member 6)"
/db_xref="GeneID:396726"
CDS 1..849
/gene="FASLG"
/gene_synonym="CD95-L; FASL; TNFSF6"
/note="CD95 ligand; tumor necrosis factor (ligand)
superfamily, member 6; fas antigen ligand"
/codon_start=1
/product="tumor necrosis factor ligand superfamily member
6"
/protein_id="NP_998971.1"
/db_xref="GeneID:396726"
/translation="MQQPFNYPYPQIFWVDSSATSPWASPGSVFPCPASVPGRPGQRR
PPPPPPPPPPPPTLLPSRPLPPLPPPSLKKKRDHNAGLCLLVMFFMVLVALVGLGLGM
FQLFHLQKELTELRESASQRHTESSLEKQIGHPNLPSEKKELRKVAHLTGKPNSRSIP
LEWEDTYGIALVSGVKYMKGSLVINDTGLYFVYSKVYFRGQYCNNQPLSHKVYTRNSR
YPQDLVLMEGKMMNYCTTGQMWARSSYLGAVFNLTSADHLYVNVSELSLVNFEESKTF
FGLYKL"
mat_peptide 1..390
/gene="FASLG"
/gene_synonym="CD95-L; FASL; TNFSF6"
/product="ADAM10-processed FasL form. {ECO:0000250}"
/experiment="experimental evidence, no additional details
recorded"
/note="propagated from UniProtKB/Swiss-Prot (Q9BEA8.1)"
mat_peptide 1..249
/gene="FASLG"
/gene_synonym="CD95-L; FASL; TNFSF6"
/product="FasL intracellular domain. {ECO:0000250}"
/experiment="experimental evidence, no additional details
recorded"
/note="propagated from UniProtKB/Swiss-Prot (Q9BEA8.1)"
misc_feature 244..249
/gene="FASLG"
/gene_synonym="CD95-L; FASL; TNFSF6"
/experiment="experimental evidence, no additional details
recorded"
/note="Cleavage, by SPPL2A. {ECO:0000250}; propagated from
UniProtKB/Swiss-Prot (Q9BEA8.1); cleavage site"
misc_feature 247..309
/gene="FASLG"
/gene_synonym="CD95-L; FASL; TNFSF6"
/experiment="experimental evidence, no additional details
recorded"
/note="propagated from UniProtKB/Swiss-Prot (Q9BEA8.1);
transmembrane region"
misc_feature 388..393
/gene="FASLG"
/gene_synonym="CD95-L; FASL; TNFSF6"
/experiment="experimental evidence, no additional details
recorded"
/note="Cleavage, by ADAM10. {ECO:0000250}; propagated from
UniProtKB/Swiss-Prot (Q9BEA8.1); cleavage site"
mat_peptide 391..846
/gene="FASLG"
/gene_synonym="CD95-L; FASL; TNFSF6"
/product="Tumor necrosis factor ligand superfamily member
6, soluble form. {ECO:0000250}"
/experiment="experimental evidence, no additional details
recorded"
/note="propagated from UniProtKB/Swiss-Prot (Q9BEA8.1)"
misc_feature 553..555
/gene="FASLG"
/gene_synonym="CD95-L; FASL; TNFSF6"
/experiment="experimental evidence, no additional details
recorded"
/note="N-linked (GlcNAc...) asparagine. {ECO:0000255};
propagated from UniProtKB/Swiss-Prot (Q9BEA8.1);
glycosylation site"
misc_feature 751..753
/gene="FASLG"
/gene_synonym="CD95-L; FASL; TNFSF6"
/experiment="experimental evidence, no additional details
recorded"
/note="N-linked (GlcNAc...) asparagine. {ECO:0000255};
propagated from UniProtKB/Swiss-Prot (Q9BEA8.1);
glycosylation site"
misc_feature 781..783
/gene="FASLG"
/gene_synonym="CD95-L; FASL; TNFSF6"
/experiment="experimental evidence, no additional details
recorded"
/note="N-linked (GlcNAc...) asparagine. {ECO:0000255};
propagated from UniProtKB/Swiss-Prot (Q9BEA8.1);
glycosylation site"
exon 1..351
/gene="FASLG"
/gene_synonym="CD95-L; FASL; TNFSF6"
/inference="alignment:Splign:2.1.0"
exon 352..397
/gene="FASLG"
/gene_synonym="CD95-L; FASL; TNFSF6"
/inference="alignment:Splign:2.1.0"
exon 398..454
/gene="FASLG"
/gene_synonym="CD95-L; FASL; TNFSF6"
/inference="alignment:Splign:2.1.0"
exon 455..849
/gene="FASLG"
/gene_synonym="CD95-L; FASL; TNFSF6"
/inference="alignment:Splign:2.1.0"
ORIGIN
1 atgcagcagc ccttcaatta cccatacccc caaatcttct gggtggacag cagtgctacc
61 tctccctggg cctccccagg ctcagtcttc ccctgtccag cttctgtgcc aggaaggcca
121 gggcaaagga ggccaccacc accaccgccg ccaccgccac caccaccaac actcctgcca
181 tcaagaccgc tgcctccact gccaccgcca tctctgaaga agaagaggga ccacaatgca
241 ggcctgtgtc tccttgtgat gttcttcatg gttctggtgg ccctggttgg attggggctg
301 gggatgtttc agctcttcca cctacagaag gagctgactg aactcagaga gtctgccagc
361 caaaggcata cagaatcatc tttggagaag caaataggtc accccaatct accctctgag
421 aaaaaggagc tgagaaaagt ggcccactta acaggcaagc ctaactcaag atccatccct
481 ctggaatggg aagacaccta tggaattgcc ttggtctctg gggtgaagta tatgaagggc
541 agccttgtga tcaatgacac tgggctgtat tttgtgtatt ccaaagtgta cttccggggt
601 cagtactgca acaaccagcc cctgagtcac aaggtataca caaggaactc taggtatccc
661 caggacctgg tgctgatgga gggaaagatg atgaactatt gcactactgg ccaaatgtgg
721 gcccgcagca gctacctggg ggctgtgttc aatctcacca gcgctgacca tttatatgtc
781 aacgtatctg agctctctct ggtcaatttt gaggaatcta agacattttt tggcttatat
841 aagctctga
//
fasta格式是一种基于文本用于表示核酸序列或多肽序列的格式。其中核酸或氨基酸均以单个字母来表示,且允许在序列前添加序列名及注释。该格式已成为生物信息学领域的一项标准。
例如:
>NM_213806
ATGCAGCAGCCCTTCAATTACCCATACCCCCAAATCTTCTGGGTGGACAGCAGTGCTACC
TCTCCCTGGGCCTCCCCAGGCTCAGTCTTCCCCTGTCCAGCTTCTGTGCCAGGAAGGCCA
GGGCAAAGGAGGCCACCACCACCACCGCCGCCACCGCCACCACCACCAACACTCCTGCCA
TCAAGACCGCTGCCTCCACTGCCACCGCCATCTCTGAAGAAGAAGAGGGACCACAATGCA
GGCCTGTGTCTCCTTGTGATGTTCTTCATGGTTCTGGTGGCCCTGGTTGGATTGGGGCTG
GGGATGTTTCAGCTCTTCCACCTACAGAAGGAGCTGACTGAACTCAGAGAGTCTGCCAGC
CAAAGGCATACAGAATCATCTTTGGAGAAGCAAATAGGTCACCCCAATCTACCCTCTGAG
AAAAAGGAGCTGAGAAAAGTGGCCCACTTAACAGGCAAGCCTAACTCAAGATCCATCCCT
CTGGAATGGGAAGACACCTATGGAATTGCCTTGGTCTCTGGGGTGAAGTATATGAAGGGC
AGCCTTGTGATCAATGACACTGGGCTGTATTTTGTGTATTCCAAAGTGTACTTCCGGGGT
CAGTACTGCAACAACCAGCCCCTGAGTCACAAGGTATACACAAGGAACTCTAGGTATCCC
CAGGACCTGGTGCTGATGGAGGGAAAGATGATGAACTATTGCACTACTGGCCAAATGTGG
GCCCGCAGCAGCTACCTGGGGGCTGTGTTCAATCTCACCAGCGCTGACCATTTATATGTC
AACGTATCTGAGCTCTCTCTGGTCAATTTTGAGGAATCTAAGACATTTTTTGGCTTATAT
AAGCTCTGA
处理步骤
- 将文件夹下gb文件批量读取
- 将各个gb文件中的登录号和具体序列抽提出来,并写入fasta文件
- 将fasta文件进一步处理去掉换行符,使一个完整的序列中没有换行符
- 将所有处理好的fasta文件存入一个新建的子文件夹中
Python脚本代码如下:
#!/usr/bin/env python
# -*- encoding: utf-8 -*-
'''
@File : gb2fasta.py
@Time : 2020/07/04 14:15:13
@Author : Ai
@Version : 1.0
@Contact : aqy0716@163.com
@License : (C)Copyright 2020 SCAU
@Desc : 将gb文件转换为fasta文件,同时转成无换行符,最后存入新的子文件夹中
'''
# here put the import lib
import os
import shutil
def gb2fasta(path):
# 从gb文件中抽取登录号和具体序列信息存入同名fasta文件
# 读取文件夹相关信息
for root,dirs,files in os.walk(path):
for file in files:
# 打印文件所属目录
print(root + '\\' + file)
# 获取文件路径
path_gb = os.path.join(root,file)
flag=0
if path_gb[-2:] == 'gb':
# 打开新建fasta文件准备写入
fasta = open (path_gb[:-2]+'fasta','w')
# 打开gb文件,准备读取序列信息并写入fasta文件
with open (path_gb,'r') as f:
# 逐行扫描
for line in f:
# 如果是ACCESSION行,则写入fasta文件作为序列标题
if line[0:9] =='ACCESSION':
fasta.writelines('>'+line.split()[1]+'\n')
# 如果是ORGIN行,代表是序列
elif line[0:6] =='ORIGIN':
flag = 1
elif flag == 1:
#通过空格符(空格 换行 制表)对字符串进行切片
s = line.split()
#非空切片字符打印
if s != []:
#print(s)
#去掉列表首个元素(数字序号)后,连接所有元素即为完整序列按行写入fasta文件
seq=''.join(s[1:])
fasta.writelines(seq.upper()+'\n')
fasta.close()
def multi2single(path):
#此函数功能为: 将多行序列转换为单行序列(即去掉换行符),成为标准fasta文件
for root,dirs,files in os.walk(path):
for file in files:
path_full = os.path.join(root,file)
# 有fasta 且不为 single.fasta,才进行单行转换,否则会重复创建文件夹
a = path_full[-5:] == 'fasta'
if path_full[-12:] == 'single.fasta':
b = True
else:
b = False
b = bool(1-b)
if a & b :
fr=open(path_full, 'r')
fw=open(path_full[:-6]+'_single.fasta', 'w')
seq={}
for line in fr:
if line.startswith('>'): #判断字符串是否以‘>开始'
name=line.split()[0] #以空格为分隔符,并取序列为0的项。
seq[name]=''
else:
seq[name]+=line.replace('\n', '')
fr.close()
for i in seq.keys():
fw.write(i)
fw.write('\n')
fw.write(seq[i])
fw.write('\n')
fw.close()
def copy2subdir(path):
# 将生成的_single.fasta文件存入新的子文件夹.\singl_fasta\中
root1 = path
for root,dirs,files in os.walk(path):
subdir = os.path.join(root1,'singl_fasta\\')
for file in files:
oldfile = os.path.join(root1,str(file))
newfile = os.path.join(root1,'singl_fasta\\',str(file))
if not os.path.exists(subdir):
os.makedirs(subdir)
print("目录创建成功!")
if oldfile[-12:] == 'single.fasta':
if not os.path.exists(newfile):
shutil.copyfile(oldfile , newfile)
print("\n你的fasta文件保存在: " + subdir + "文件夹下\n")
if __name__ == "__main__":
path = input("请输入路径") #此处输入 D:\docu\gb2fasta
gb2fasta(path)
multi2single(path)
copy2subdir(path)
运行情况
PS D:\vscode_python_magic> & d:/ruanjiancangku/python_projectkotin/venv/Scripts/python.exe d:/vscode_python_magic/实验室-magic/序列处理/gb2fasta.py D:\docu\gb2fasta\1-FASLG-swine-849bp-NM_213806.fasta D:\docu\gb2fasta\1-FASLG-swine-849bp-NM_213806.gb ['1', 'atgcagcagc', 'ccttcaatta', 'cccatacccc', 'caaatcttct', 'gggtggacag', 'cagtgctacc'] ['61', 'tctccctggg', 'cctccccagg', 'ctcagtcttc', 'ccctgtccag', 'cttctgtgcc', 'aggaaggcca'] ['121', 'gggcaaagga', 'ggccaccacc', 'accaccgccg', 'ccaccgccac', 'caccaccaac', 'actcctgcca'] ['181', 'tcaagaccgc', 'tgcctccact', 'gccaccgcca', 'tctctgaaga', 'agaagaggga', 'ccacaatgca'] ['241', 'ggcctgtgtc', 'tccttgtgat', 'gttcttcatg', 'gttctggtgg', 'ccctggttgg', 'attggggctg'] ['301', 'gggatgtttc', 'agctcttcca', 'cctacagaag', 'gagctgactg', 'aactcagaga', 'gtctgccagc'] ['361', 'caaaggcata', 'cagaatcatc', 'tttggagaag', 'caaataggtc', 'accccaatct', 'accctctgag'] ['421', 'aaaaaggagc', 'tgagaaaagt', 'ggcccactta', 'acaggcaagc', 'ctaactcaag', 'atccatccct'] ['481', 'ctggaatggg', 'aagacaccta', 'tggaattgcc', 'ttggtctctg', 'gggtgaagta', 'tatgaagggc'] ['541', 'agccttgtga', 'tcaatgacac', 'tgggctgtat', 'tttgtgtatt', 'ccaaagtgta', 'cttccggggt'] ['601', 'cagtactgca', 'acaaccagcc', 'cctgagtcac', 'aaggtataca', 'caaggaactc', 'taggtatccc'] ['661', 'caggacctgg', 'tgctgatgga', 'gggaaagatg', 'atgaactatt', 'gcactactgg', 'ccaaatgtgg'] ['721', 'gcccgcagca', 'gctacctggg', 'ggctgtgttc', 'aatctcacca', 'gcgctgacca', 'tttatatgtc'] ['781', 'aacgtatctg', 'agctctctct', 'ggtcaatttt', 'gaggaatcta', 'agacattttt', 'tggcttatat'] ['841', 'aagctctga'] ['//'] D:\docu\gb2fasta\2-LTA-swine-1584bp-NM_214453.fasta D:\docu\gb2fasta\2-LTA-swine-1584bp-NM_214453.gb ['1', 'agaaaggggc', 'ccacaggggt', 'cccgcacagc', 'aggtgagact', 'ctcccacccc', 'atctcctagg'] ['61', 'gctgtccggg', 'tgctggactc', 'ccccctcact', 'tcggtccctc', 'cgcccgctcc', 'ctggccttcc'] ['121', 'tgcccctcct', 'gcatcttcac', 'cccggcctgg', 'gccttggtgg', 'gtttggtttt', 'ggtttgttct'] ['181', 'ctctgattct', 'ttatctgtca', 'ggctctttct', 'agctctcaca', 'cactctgatc', 'cctctctgtt'] ['241', 'cccttcccat', 'ctctgtttct', 'ctctgggtct', 'ccccctgctc', 'acctcgggat', 'ttccctgagt'] ['301', 'gcctctggtc', 'cccttctctg', 'tctggcgccc', 'cgtctcttgt', 'ctctcggggt', 'ggctgtctcc'] ['361', 'gagggcagga', 'ggccttcttc', 'cgcaggtgcc', 'ccgccccgct', 'cactgtctct', 'ctccccccac'] ['421', 'aggttttccc', 'catgacacca', 'cctggacgcc', 'tctacctccg', 'gagggtgtgc', 'agcaccccca'] ['481', 'tcctcctcct', 'cctggggctg', 'ctgctggccc', 'tgccgcccga', 'ggcccagggg', 'ctccctggcg'] ['541', 'tcggcctccc', 'accctcagct', 'gcacagcctg', 'cccatcagca', 'ccccccaaag', 'cacttggcca'] ['601', 'gaggcaccct', 'caaacctgcc', 'gctcacctcg', 'ttggagaccc', 'cagcaccccg', 'gactcactgc'] ['661', 'gctggagagc', 'gaacacggat', 'cgtgccttcc', 'tccgccatgg', 'cttcttgctg', 'agcaacaact'] ['721', 'ccctgctggt', 'ccccaccagt', 'ggcctctact', 'ttgtctactc', 'ccaggtcgtc', 'ttctccgggg'] ['781', 'aaggctgctt', 'ccccaaggcc', 'acccccaccc', 'ctctctacct', 'ggcccacgag', 'gtccagctct'] ['841', 'tctcctccca', 'gtaccccttc', 'cacgtgccgc', 'tcctcagcgc', 'tcagaagtcc', 'gtgtgccccg'] ['901', 'ggccacaggg', 'accttgggtg', 'cgctctgtgt', 'accagggggc', 'tgtgttcctg', 'ctcacccagg'] ['961', 'gagatcagct', 'gtccacacac', 'acagacggca', 'ccccccacct', 'gctcctcagc', 'cccagtagcg'] ['1021', 'tcttctttgg', 'agccttcgct', 'ctatagaaga', 'atccagaaag', 'aaaaaaattg', 'gtttcaaggc'] ['1081', 'cttctcccct', 'tttcacctcc', 'cttatgacca', 'cttcggaggt', 'caccgcgcct', 'ctcctctgac'] ['1141', 'aatttccaac', 'agtctcatct', 'tcccccacgc', 'tcagcacctg', 'gagcttctgt', 'agaaggaatt'] ['1201', 'ctaggcacct', 'cgggggaact', 'ggaaccaccc', 'cggatgctct', 'gctgaggatc', 'tgaatgcccg'] ['1261', 'cctggagccc', 'ttcccctgtc', 'ctgcccgtct', 'aggggccctc', 'gtccaggacg', 'tggaagggaa'] ['1321', 'gctgacccat', 'gagggacttt', 'gaacggatga', 'ccggagcggt', 'gtgggggggt', 'tatttatgaa'] ['1381', 'ggggaaaatt', 'aaattattta', 'tttatggagg', 'atggagagaa', 'gggaatcaca', 'gagggatgtc'] ['1441', 'agaagagtgt', 'gacacatgtg', 'cccaagagat', 'aaagtgacag', 'aaggcatggg', 'ctccagatga'] ['1501', 'cccggccaga', 'gagggcaaag', 'tggctcagga', 'aggggctgct', 'tgactggagg', 'ctcatgagga'] ['1561', 'gacggctgac', 'cctcgatgaa', 'accc'] ['//'] D:\docu\gb2fasta\3-LTB-swine-950bp-NM_001185138.fasta D:\docu\gb2fasta\3-LTB-swine-950bp-NM_001185138.gb ['1', 'tcggatgggg', 'gcaccggggc', 'tggagggccg', 'gggtaggagg', 'ccccagggga', 'agggatgcct'] ['61', 'cctgctggcc', 'gtggcagggg', 'ccacttccct', 'ggtgaccctc', 'ctgctggccg', 'tgcctatcac'] ['121', 'ggtcctggct', 'gtgctggcct', 'tggtgcccca', 'ggagcaggga', 'gaactggtaa', 'cagggaccgc'] ['181', 'tgacccaggc', 'acccaggcgg', 'aggcccagca', 'gcgattggag', 'tccaaggaga', 'cgccagagga'] ['241', 'ggaggcagaa', 'acagatctca', 'gccccaggct', 'cccagctgcc', 'cacctcattg', 'gcgcttggat'] ['301', 'cacgggtcag', 'gggctaggct', 'gggaggcgaa', 'gaaagaagag', 'gcgtttctga', 'ggagcgggac'] ['361', 'gcagttctct', 'ggcgcggagg', 'gcctggccct', 'cccgcaggac', 'ggcctctact', 'acctctactg'] ['421', 'tcacgtcggc', 'taccggggcc', 'gggcacctcc', 'tcccggcggg', 'gaccccctgg', 'accgctcggt'] ['481', 'cacgctgctc', 'agccggctgt', 'accgggcggg', 'gggcgcctac', 'ggaccgggga', 'ctcccgagct'] ['541', 'gctgctggag', 'ggcgcggaga', 'ctgtgactcc', 'ggtcttggac', 'cccagtcgga', 'ggcacgagta'] ['601', 'cgggcccctc', 'tggtacacga', 'gcgtggggtt', 'cggtggcctg', 'gtgcagctcc', 'ggaggggcga'] ['661', 'gagggtgtac', 'gttaatatca', 'gtcaccccga', 'tatggtggat', 'tacaggagag', 'gaaagacctt'] ['721', 'cttcggggcg', 'gtgatggtgg', 'gctgaggact', 'gtccgcggcc', 'cgagaggacc', 'actgcatggt'] ['781', 'gggagtgtgt', 'cgatggatca', 'agcccagaca', 'cggggtccca', 'gacaccaggc', 'cagacaccat'] ['841', 'ggccgtgggg', 'aaaatgcagg', 'agatcgtgtg', 'gaaaactgat', 'tttgagcctg', 'atgaaaataa'] ['901', 'agaatgtaaa', 'agctttaatn', 'gctgcccatg', 'ccaaaaaaaa', 'aaaaaaaaaa'] ['//'] D:\docu\gb2fasta\4-TNF-swine-1666bp-NM_214022.fasta D:\docu\gb2fasta\4-TNF-swine-1666bp-NM_214022.gb ['1', 'cccagagtga', 'ggacaccagg', 'ggaccagcca', 'ggagagagac', 'aagccatctc', 'caggaccccc'] ['61', 'tagaaataac', 'ctctcagaag', 'acacaccccc', 'gaacaggcag', 'ccggacgact', 'ctctccctct'] ['121', 'cacacgctgc', 'cccggggcgc', 'caccatctcc', 'cagctggacc', 'tgagcccctc', 'tgaaaaagac'] ['181', 'accatgagca', 'ctgagagcat', 'gatccgagac', 'gtggagctgg', 'cggaggaggc', 'gctcgccaag'] ['241', 'aaggccgggg', 'gcccccaggg', 'ctccaggagg', 'tgcctgtgcc', 'tcagcctctt', 'ctccttcctc'] ['301', 'ctggtcgcag', 'gagccaccac', 'gctcttctgc', 'ctactgcact', 'tcgaggttat', 'cggcccccag'] ['361', 'aaggaagagt', 'ttccagctgg', 'ccccttgagc', 'atcaaccctc', 'tggcccaagg', 'actcagatca'] ['421', 'tcgtctcaaa', 'cctcagataa', 'gcccgtcgcc', 'cacgttgtag', 'ccaatgtcaa', 'agccgaggga'] ['481', 'cagctccaat', 'ggcagagtgg', 'gtatgccaat', 'gccctcctgg', 'ccaacggcgt', 'gaagctgaaa'] ['541', 'gacaaccagc', 'tggtggtgcc', 'gacagatggg', 'ctgtacctca', 'tctactccca', 'ggtcctcttc'] ['601', 'aggggccaag', 'gctgcccttc', 'caccaacgtt', 'ttcctcactc', 'acaccatcag', 'ccgcatcgcc'] ['661', 'gtctcctacc', 'agaccaaggt', 'caacctcctc', 'tctgccatca', 'agagcccttg', 'ccagagggag'] ['721', 'acccccgagg', 'gggccgaggc', 'caagccctgg', 'tacgaaccca', 'tctacctggg', 'aggggtcttc'] ['781', 'cagctggaga', 'aggatgatcg', 'actcagtgcc', 'gagatcaacc', 'tgcccgacta', 'tctggacttt'] ['841', 'gctgaatctg', 'ggcaggtcta', 'ttttgggatc', 'attgccctgt', 'gagggggcag', 'gacatccgtt'] ['901', 'ccctcccctg', 'tccatccctt', 'tattatttta', 'ctccttcaga', 'ccccctcacg', 'tccttctggt'] ['961', 'ttagaaagag', 'aatgaggggc', 'tggggactgg', 'gctccaagct', 'taaaacttta', 'aacaacaaca'] ['1021', 'gcaacactta', 'gaaatcaggg', 'attcagggat', 'gtgtggcctg', 'gacaaccagg', 'cactgaccac'] ['1081', 'caccaagaat', 'tggaactggg', 'gcttccagac', 'tcgctggggt', 'ccttgggttt', 'ggattcctgg'] ['1141', 'atgcaacctg', 'ggacatctgg', 'aatgtggctg', 'ccagggaagc', 'ttgggttcca', 'atcggaatac'] ['1201', 'ttcagaacat', 'tccttgagaa', 'gatttcacct', 'caatcttgat', 'gactttttag', 'gcttcccttt'] ['1261', 'cttccaattt', 'tccagacttc', 'cctgggatgg', 'ggagcccagc', 'cccaaacccc', 'acaggccagc'] ['1321', 'tccctcttat', 'ttatatttgc', 'acttggcatt', 'attatttatt', 'tatttattta', 'ttatttattt'] ['1381', 'actagtgaat', 'gtatttattc', 'aggagggcga', 'ggtgtcctgg', 'gagacccagc', 'ataagggctg'] ['1441', 'ccttggttca', 'gatgtgtttt', 'ctgtgaaaac', 'ggagctgaac', 'tgtaggttgc', 'tcccacctgg'] ['1501', 'cctcctagcc', 'tctgtgcctc', 'cttttgctta', 'tgtttttaaa', 'aacaaatatt', 'tatctgatcg'] ['1561', 'agttgtctaa', 'ataatgctga', 'tttggtgact', 'aacttgtcgc', 'tacatcgctg', 'aacctctgct'] ['1621', 'ccccagggga', 'gttgtgtctg', 'taaccgccct', 'actggtcagt', 'ggcgag'] ['//'] D:\docu\gb2fasta\5-TNFSF4-swine-549bp-NM_001025217.fasta D:\docu\gb2fasta\5-TNFSF4-swine-549bp-NM_001025217.gb ['1', 'atggaagggg', 'tccaacccct', 'agatgaaaat', 'gtgggaaacg', 'caccaggacg', 'aagactcttg'] ['61', 'aggaacaagc', 'tattgttggt', 'ggcctccgta', 'attcagggtc', 'tggggttgct', 'cctgtgtctc'] ['121', 'acctacatct', 'gcctgcacct', 'ctatgctcag', 'gtgccatctc', 'agtaccctcc', 'aattcagagt'] ['181', 'atcaaagtac', 'aatttaccaa', 'gtgtgaaaat', 'gataatggtt', 'tcatcatcac', 'accctcaagc'] ['241', 'aaggatggaa', 'ccatgaaagt', 'gcaaaacaac', 'tcaatcatca', 'tcaactgtga', 'tgggttctat'] ['301', 'ctcatctccc', 'tgaagggtta', 'cttttctcag', 'gagctcagcc', 'tcatgcttca', 'gtaccggaag'] ['361', 'ggtcggaaac', 'ctctcttctc', 'cctgaacaag', 'gtcaagtctg', 'tggactctgt', 'cacagtagcc'] ['421', 'gatctggctt', 'tcaaggacaa', 'ggtcttcctg', 'aacgtgacca', 'ctcatagtgc', 'ctcctgtgaa'] ['481', 'gacattcagg', 'tgaatggtgg', 'ggaattgatt', 'ctcattcatc', 'aaaatcctgg', 'tggattctgt'] ['541', 'gtctactga'] ['//'] D:\docu\gb2fasta\6-TNFSF10-swine-1696bp-NM_001024696.fasta D:\docu\gb2fasta\6-TNFSF10-swine-1696bp-NM_001024696.gb ['1', 'agcagtcaga', 'ccctgcctgg', 'accatggcgg', 'tgatgcagac', 'tccaggaggc', 'cccagccccg'] ['61', 'ggcagacctg', 'tgtgttgatc', 'ctgatcttca', 'cagtgctcct', 'gcaagccctc', 'tgtgtggcct'] ['121', 'tgacttacgt', 'gtacttcacc', 'aatgaactga', 'aacagatgca', 'ggacaagtac', 'tccaaaagcg'] ['181', 'gtatagcttg', 'cttcttaaag', 'gaagatgaca', 'gtttctggga', 'tcccaccgat', 'gacgagagaa'] ['241', 'tgctcagccc', 'ctgctggcag', 'gtgaagtggc', 'agctacgtca', 'gtttgtgaga', 'aagatgattt'] ['301', 'tgagaaccta', 'tgaggaaacc', 'atttctacag', 'tttcagaaaa', 'gcaacaaggc', 'attcctcacc'] ['361', 'tagaaagaga', 'aaaaggtcca', 'cagagagtgg', 'ctgctcacat', 'aactggaacc', 'agtaggaaaa'] ['421', 'gaagcacatt', 'tccatctcta', 'agctccaaat', 'atgaaaaagc', 'tttgggccag', 'aaaataaact'] ['481', 'cctgggaatc', 'atcaagaaaa', 'ggacattcat', 'tcttgaataa', 'ttttcacttg', 'aggaatggag'] ['541', 'agctggttat', 'ccatcaaaca', 'gggttttact', 'acatctattc', 'ccaaacatac', 'tttcgatttc'] ['601', 'aggaacctga', 'ggaaattttg', 'ggaacggttt', 'ctacagaagg', 'gaacagaaag', 'aaaaacaggc'] ['661', 'aaatgataca', 'gtatatttac', 'aaatggacaa', 'gctatcctga', 'ccctatactg', 'ctgatgaaaa'] ['721', 'gtgctagaaa', 'tagttgttgg', 'tctaaagatt', 'cagaatatgg', 'actctattcc', 'atctatcaag'] ['781', 'gtggaatatt', 'tgagcttaag', 'gaagatgacc', 'gaatttttgt', 'ctctgttact', 'aatgagcaac'] ['841', 'tgattgacat', 'ggaccaagaa', 'gccagttttt', 'tcggggcctt', 'tttaattggc', 'taaatgatct'] ['901', 'gcagggaaaa', 'aaaccatgcc', 'ccagagtgac', 'tattcagagt', 'cgtatactgt', 'gaaaatattc'] ['961', 'cagcagagcc', 'aataggttaa', 'ggcagcctga', 'gcaaagaggc', 'ctcaacccaa', 'aggctcaaca'] ['1021', 'acacaagctt', 'tttggaaagt', 'gaaaagtgac', 'caattccttc', 'caggaaaatg', 'aaactgccaa'] ['1081', 'gagaccttgt', 'ggagctctgc', 'ctgatgtcat', 'tttgctagta', 'aacatctaga', 'agatactctg'] ['1141', 'tctccaaatt', 'tgtgtaacaa', 'ttaacacctc', 'ctgcctttat', 'catctaatcc', 'tgtgaagatt'] ['1201', 'ctagaagaaa', 'gagtagtgat', 'ccatctcagg', 'tgggaataag', 'ggacaacatt', 'cccaaaacta'] ['1261', 'aagagaaaag', 'ggcagcactg', 'aaaggtcaca', 'gtcaatatat', 'gcagtttcag', 'tacaaacata'] ['1321', 'acaaattaaa', 'gctacgttta', 'gtggacaagg', 'agctacttct', 'gaatggtttg', 'tgcttttctc'] ['1381', 'tactaaaaat', 'caggctggcc', 'aaaagcactc', 'agggtatttt', 'tgataaagga', 'ctctaaaata'] ['1441', 'agtgataaag', 'tatggcgata', 'cctcagaaaa', 'ctaaatacag', 'aactaccaca', 'tgacccagca'] ['1501', 'atcccactcc', 'tgtgcatata', 'tctggacaaa', 'actttccttg', 'aaaaagatac', 'attcatctct'] ['1561', 'atgctcattg', 'cagcactatt', 'cacagtagcc', 'aagacatgga', 'aacacctata', 'tgtctatgaa'] ['1621', 'tggatgaata', 'gattaagaag', 'gtgtgttatg', 'tatacatant', 'ggaatactgg', 'gaagccataa'] ['1681', 'aaaggacaaa', 'gaggcc'] ['//'] D:\docu\gb2fasta\7-TNFSF13b-swine-1013bp-NM_001097498.fasta D:\docu\gb2fasta\7-TNFSF13b-swine-1013bp-NM_001097498.gb ['1', 'tctaggaggg', 'aaatggatga', 'ctccacgggg', 'gagcagtcac', 'gcctttcttg', 'ccttagcacg'] ['61', 'agagaagaaa', 'tgaaactgaa', 'ggagacggtc', 'cccatcctcc', 'cccagaagga', 'aagcccctct'] ['121', 'gtccgcatct', 'ccaaagatgg', 'gaagctgctg', 'gtcgtgaccc', 'tgctgctggc', 'cctgctgtcc'] ['181', 'tgctgcctca', 'cggggatctt', 'tgcaccacca', 'gctccaaggg', 'agagcagctc', 'cattcaaagc'] ['241', 'aacaggagta', 'agcgcgccgc', 'gcaggatgcg', 'gaggagacag', 'tcactcagga', 'ctgcttgcaa'] ['301', 'ttgattgcag', 'acagtgacat', 'gcctactata', 'cgaaaaggag', 'cttatacatt', 'tgttccatgg'] ['361', 'cttctcagct', 'ttaaaagagg', 'aagagcccta', 'gaagaaaaag', 'aaaataaaat', 'cgtggtcaaa'] ['421', 'gaaacgggtt', 'acttttttat', 'atacggtcag', 'gttttataca', 'ccgataacac', 'ctttgccatg'] ['481', 'gggcatctca', 'tacagaggaa', 'gaaagtccat', 'gtctttgggg', 'atgaactgag', 'tctggtgact'] ['541', 'ttgttccgat', 'gtattcaaaa', 'tatgcctgaa', 'acactaccca', 'ataattcctg', 'ttattcagct'] ['601', 'ggcattgcaa', 'agctggagga', 'aggagatgaa', 'ctccaactgg', 'caataccacg', 'tgaagacgct'] ['661', 'aaaatatcac', 'gggatggaga', 'cggcacattt', 'tttggtgcat', 'tgaaacttct', 'gtgacctact'] ['721', 'tacaccttgt', 'ttgtggctct', 'tgccctccct', 'ccctctgtac', 'ctctaaagag', 'aaaacactta'] ['781', 'actggaaata', 'ccaaaagggg', 'aaaaaaaagt', 'agttaccata', 'gccttttctg', 'tgagctgttt'] ['841', 'gttttggttt', 'gctgaaacta', 'gaccaaaaca', 'ggaaatttaa', 'cagacaacca', 'cagccaaagg'] ['901', 'gtatcatgtg', 'aattacaaga', 'aatagagccc', 'atttaagaaa', 'aaatagaatt', 'agaaagactt'] ['961', 'ttcactgtaa', 'tgccatgttg', 'aacagcttag', 'tcatagcttc', 'ttgtcttgga', 'gga'] ['//'] D:\docu\gb2fasta\8-TNFSF18-swine-5005bp-XM_005667782.fasta D:\docu\gb2fasta\8-TNFSF18-swine-5005bp-XM_005667782.gb ['1', 'gcattccatt', 'taacaaatga', 'aaaggctaag', 'gcataaagaa', 'ccaggagaga', 'gaaccggaga'] ['61', 'tttcctcaat', 'tttagtgtag', 'taaatcaaca', 'gttttagtgc', 'taggaagttt', 'tttggaatag'] ['121', 'agtgtaaact', 'cagatggtag', 'gacagggtgc', 'atgaaaatat', 'ccttttctat', 'gataacttta'] ['181', 'tttctgtctg', 'atgtcagcat', 'tgaaatttca', 'gagtattaaa', 'atggtgaggt', 'atgaaaaaac'] ['241', 'taagcttgtt', 'gttattgaca', 'tatttttaaa', 'aataaaattc', 'tagtaataca', 'ttactgtttt'] ['301', 'ctagagatta', 'tctcagaatg', 'gacacattgt', 'ataccctagc', 'aattgatgaa', 'aatatttttc'] ['361', 'ccaaaccatg', 'aaccccagca', 'tccttagctc', 'cctgacctac', 'tgcctcccac', 'aaacatgata'] ['421', 'tttggagtat', 'gatagacctt', 'catcttgaat', 'ttcattcttt', 'ttcttacaaa', 'agtaattttc'] ['481', 'ttatctggaa', 'ataatttgta', 'atgttgaata', 'gttccatagt', 'tcctcttgct', 'tcagaaaata'] ['541', 'tttatttttc', 'ctcttaactt', 'cccttgttgg', 'gttttttttt', 'tttaaatcat', 'ctgtgtttgt'] ['601', 'gttggcttct', 'agccctcagt', 'tccagcacct', 'ttggtctggt', 'gccaaatgtt', 'agtcagcact'] ['661', 'taggctaaaa', 'gtatcgtttt', 'ccaacaccca', 'gatcagaagg', 'aaaactccgc', 'ctcttacacc'] ['721', 'cactacttag', 'tgctatacta', 'caaaactgac', 'tagttgaatc', 'atgtgctcat', 'tacttctgaa'] ['781', 'tttctgcttt', 'tcacaactct', 'cattcctgca', 'gagaatgagt', 'ttgagccaca', 'tggagaatat'] ['841', 'gcctttaagc', 'cattcaagtc', 'ctcacgcagc', 'acagagacca', 'tcctggaagc', 'aatggctact'] ['901', 'ctactcaaca', 'atagttattt', 'tgctattact', 'ttgctccttc', 'agtgcactaa', 'tcttaacttt'] ['961', 'tctcccactc', 'aagacctcca', 'acaggccatg', 'tgtagcgaag', 'tttggaccat', 'taccttcaaa'] ['1021', 'atggcaaatg', 'ccatctcctg', 'agccttcttg', 'tgtgaataag', 'acagatgatt', 'ggaggctgaa'] ['1081', 'gatacttcag', 'aatggcttgt', 'atttaattta', 'tggccaagtg', 'gctcccaaca', 'cagcttacaa'] ['1141', 'ggggcaagct', 'ccttttgagg', 'tgttgctacg', 'taggaatgaa', 'gaccccatac', 'aatctctaac'] ['1201', 'gaacaattct', 'acagtccaga', 'atgtaggagg', 'ggcttatgaa', 'tttcatgctg', 'gagatgtaat'] ['1261', 'agacttgata', 'ttcaatgctg', 'aacatcaggt', 'tctaaaaaat', 'aatacatact', 'gggggatctt'] ['1321', 'tctgctagca', 'aatccccaat', 'tcatctccta', 'gagactcagt', 'taggtctcct', 'catcttcagc'] ['1381', 'acatgcagag', 'atgccagtgc', 'ataggatgga', 'gaaggaagat', 'tttcaacaca', 'tacagttcat'] ['1441', 'ctgggtatac', 'aaatcaacat', 'gaacagatct', 'cctctgcatg', 'tgaagcttca', 'tttctcctgc'] ['1501', 'ttattgaatg', 'agactcagaa', 'agcactgaag', 'acatttggtt', 'acccctgatg', 'ttgggtcagc'] ['1561', 'aaagacactt', 'tactagttca', 'tgataaaatg', 'aaaatgggtg', 'gctggaagac', 'aaaatctttt'] ['1621', 'caaagtgtct', 'gtctaatcct', 'tgaacccctg', 'agtggaaaaa', 'tgaggtctat', 'tcccataata'] ['1681', 'gccttatata', 'gcatgcaaaa', 'aaagaccagg', 'gcagtagcct', 'ggtcttgttc', 'ttatattctt'] ['1741', 'ggactgtgga', 'ctgtttcaat', 'tcattcttcc', 'catattctca', 'tcttaggaga', 'cactcttaat'] ['1801', 'aaaatgtagt', 'cagagtgggt', 'gtgtggccag', 'caacactcca', 'ttttggagtt', 'gatgagatta'] ['1861', 'ggggatagag', 'aacactctta', 'ggaaatattg', 'ggacagaatt', 'tcagttggca', 'ttgaaatgga'] ['1921', 'atgcacttta', 'ttcgggaatt', 'tcacttgatt', 'tcatcatcaa', 'gtgcagggtg', 'ctctataaaa'] ['1981', 'cctgctggtc', 'aaaaggctag', 'ctttcaatct', 'tcacatagca', 'gttcatgaga', 'atttactggt'] ['2041', 'gtatgtatct', 'aaccatgtca', 'atgacaaaga', 'gtaatcatta', 'gtagtaagat', 'ctaacccccc'] ['2101', 'aaattggtat', 'taccagtact', 'gtactttgca', 'actgtgcaga', 'gccagctaaa', 'aatatgaaat'] ['2161', 'cattacatga', 'caaagcactt', 'tcatatacca', 'catggcaact', 'cgatagattt', 'aatggggcag'] ['2221', 'atatttttgg', 'cacaatttta', 'actatgagaa', 'tacagaggca', 'gatggaatag', 'aggtaacttg'] ['2281', 'gttcagttca', 'tagaactagt', 'aattaacagg', 'cacctgggct', 'tcccactgta', 'ttatactata'] ['2341', 'ttagctttac', 'gattggtatt', 'tctgctatca', 'tgttagaagc', 'ctataaactt', 'taacagattt'] ['2401', 'aaaattttca', 'gacagtatat', 'tcccttttag', 'tccaacagca', 'attttttcct', 'ttctcagcaa'] ['2461', 'atttcttttc', 'ttttctttgc', 'ctggagcagg', 'gtacccaggg', 'tgttattcaa', 'gacttactac'] ['2521', 'aacttaatct', 'ccttccttac', 'tttggtcaaa', 'tgtgttaact', 'tccaaaaata', 'atgaataata'] ['2581', 'ctcaattcag', 'ggacagtctg', 'ttaaattttt', 'ggactctgca', 'aaattaacta', 'gctgcttatg'] ['2641', 'ggttgttatt', 'aaaaggtatg', 'taggtaatgt', 'gattacatga', 'aaacccaatt', 'taaaatattt'] ['2701', 'atggatattt', 'gtaaaaaatc', 'tacattatgt', 'taattaatag', 'tatcaccatt', 'aaaaactaat'] ['2761', 'ttaagaatat', 'ttgtattgta', 'tgtaagaaaa', 'actgcttgga', 'agcagactaa', 'gcctgaggcc'] ['2821', 'aagatgcctc', 'atagtatgtc', 'tttttttttt', 'tttttttaaa', 'tacatctgct', 'gagcagctgt'] ['2881', 'agggacaaag', 'actggggtac', 'ctggttcctc', 'ttgtatttgt', 'gtatcatctc', 'aggaaattaa'] ['2941', 'agttacataa', 'catacatata', 'tttatggaaa', 'cgtggtattg', 'atgttaactt', 'ataagcagta'] ['3001', 'gtgtgctgga', 'gtgggctagc', 'actagctcag', 'gagagctgtt', 'aaatttttat', 'taattgtgta'] ['3061', 'gtctggttat', 'taaatcatta', 'tccttgaaat', 'tggccatggt', 'aggacaattt', 'ataccatgtg'] ['3121', 'aattagcaaa', 'tgctacaaat', 'cagggctttt', 'ctttttggaa', 'agcccatgca', 'ccagcacacc'] ['3181', 'actgtttata', 'aaactcttct', 'taatgactcc', 'tctcagcccc', 'tgcctcagta', 'ttacaacagt'] ['3241', 'caaggcaggc', 'aaggaaagtg', 'tcttactctc', 'agcaaaagcc', 'ccacagataa', 'atcatttctc'] ['3301', 'agggcaggtg', 'gaggaatcta', 'cagctgtaac', 'cagatagata', 'gctaccaaca', 'tatgaccttt'] ['3361', 'gaatttccct', 'agtgttgaaa', 'tttcaggctt', 'tgttttcaat', 'gtatactctg', 'ttcccttgtt'] ['3421', 'tcttcaaaac', 'agtgtttata', 'ttttaaactg', 'acaataaaat', 'gtttgtacat', 'gggctgtagc'] ['3481', 'tgatttatct', 'atgggttatc', 'tgacaagagt', 'tagtgatcta', 'gagaatgtgc', 'atcagagaaa'] ['3541', 'tccttgtcca', 'tatgaaacac', 'tgctgagaga', 'gtgagaggca', 'aaagtcagag', 'ctgggggttt'] ['3601', 'tgtcccaatt', 'tctgatgggt', 'taggcaaata', 'atttcactag', 'ctgactttca', 'cccacctcct'] ['3661', 'caactctacc', 'ccacccatcc', 'accctcacca', 'tcaaagccat', 'catcaaccct', 'ctcatactaa'] ['3721', 'ttgtttttac', 'cttctatttt', 'gcctcatcaa', 'agctgacctg', 'gagtgctttt', 'tatcactgtg'] ['3781', 'ggttaagtct', 'ctacttctga', 'agatgacagt', 'taattgatca', 'cagtcctgtg', 'tatttgatgg'] ['3841', 'aacagataaa', 'ctccttagat', 'gcattatatc', 'tttgcaactt', 'ggagataaaa', 'ttcctgtctt'] ['3901', 'tgtgtgttct', 'tcctcaaaga', 'agcaattttc', 'attggctggg', 'ggtgctagta', 'cgttttttat'] ['3961', 'tagtgagaag', 'gagaagcgaa', 'aacccaattg', 'acttatgggc', 'tagtggtctg', 'aggcttacct'] ['4021', 'tcaagcagag', 'tttttcaaag', 'tgctatctat', 'ggactctaca', 'ttagatttgc', 'ccagtatact'] ['4081', 'tggccaaaaa', 'ccaaagaaaa', 'accccaaaga', 'taccttgacc', 'tcattcatag', 'tccactaaat'] ['4141', 'aaatgtttct', 'aggagtgagg', 'tctaggaatc', 'aaatttgtaa', 'tgagctctga', 'cataggttat'] ['4201', 'tcttatacta', 'cagtttaaaa', 'accactgctt', 'tatggaatgc', 'aattttgaat', 'ctcagcagaa'] ['4261', 'tgtttctcca', 'aatatttaaa', 'aacagtcaaa', 'gcttataaac', 'caatgaagtt', 'aaaaagcaaa'] ['4321', 'gcaaaaagca', 'aaaccaaaat', 'tggtattcat', 'gagacttacc', 'ataagccatc', 'tctcatcaca'] ['4381', 'aacaatggcc', 'ttctatcctc', 'tatttcagtt', 'tgaaaaaaag', 'gtcaggctct', 'tcactttcca'] ['4441', 'taatctaatg', 'gcaactttgc', 'cagttgaatg', 'ctggcaatcc', 'atagcatatg', 'atccttgtca'] ['4501', 'taatcagtac', 'tccattatct', 'acctgtatgc', 'tgtcaactct', 'caaatactta', 'tctctagcca'] ['4561', 'aagttgttct', 'cctttaagtc', 'ttataactcc', 'acctgtctgc', 'acagtaattt', 'cacattggac'] ['4621', 'atacaatgga', 'catctcaaat', 'tcaacttacc', 'ccaaatgaat', 'cttttaaggt', 'ttcaactaaa'] ['4681', 'gcagcttttt', 'tcgaatccct', 'cctcatttca', 'attgtatctc', 'atgagccaaa', 'aacctctgaa'] ['4741', 'tcatggagtc', 'attcctgact', 'tctatcttta', 'atctgtttaa', 'tcttaggaaa', 'actattatct'] ['4801', 'ttactttcag', 'agaatctgac', 'ctctttttgg', 'ctatggctct', 'tgtctttgga', 'ccaatatttt'] ['4861', 'tttcagtatc', 'ttcataactt', 'attacagtat', 'ctacatactt', 'attacattca', 'ggatttttgg'] ['4921', 'actagaaact', 'gaattttatc', 'aaagtcttag', 'agtttatata', 'tctctcatat', 'atgaaaaacc'] ['4981', 'acagctaaca', 'tcattcacaa', 'tggtg'] ['//'] D:\docu\gb2fasta\gb2fasta.py 目录创建成功! 你的fasta文件保存在: D:\docu\gb2fasta\singl_fasta\文件夹下
程序运行前 D:\docu\gb2fasta\ 文件夹下 文件

运行前
程序运行后 D:\docu\gb2fasta\ 文件夹下 文件
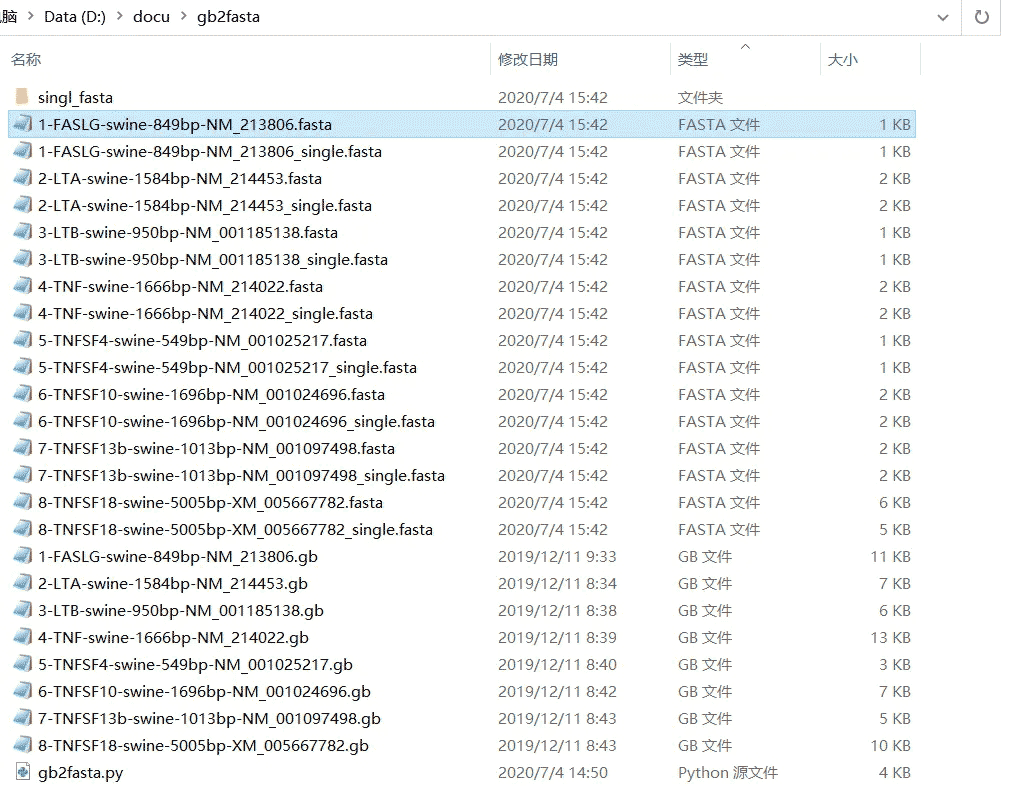
运行后
程序运行后 D:\docu\gb2fasta\ singl_fasta\ 文件夹下 文件

以上就是Python实现GB格式序列文件转换Fasta格式文件的详细内容,更多关于Python GB转换Fasta格式文件的资料请关注我们其它相关文章!

